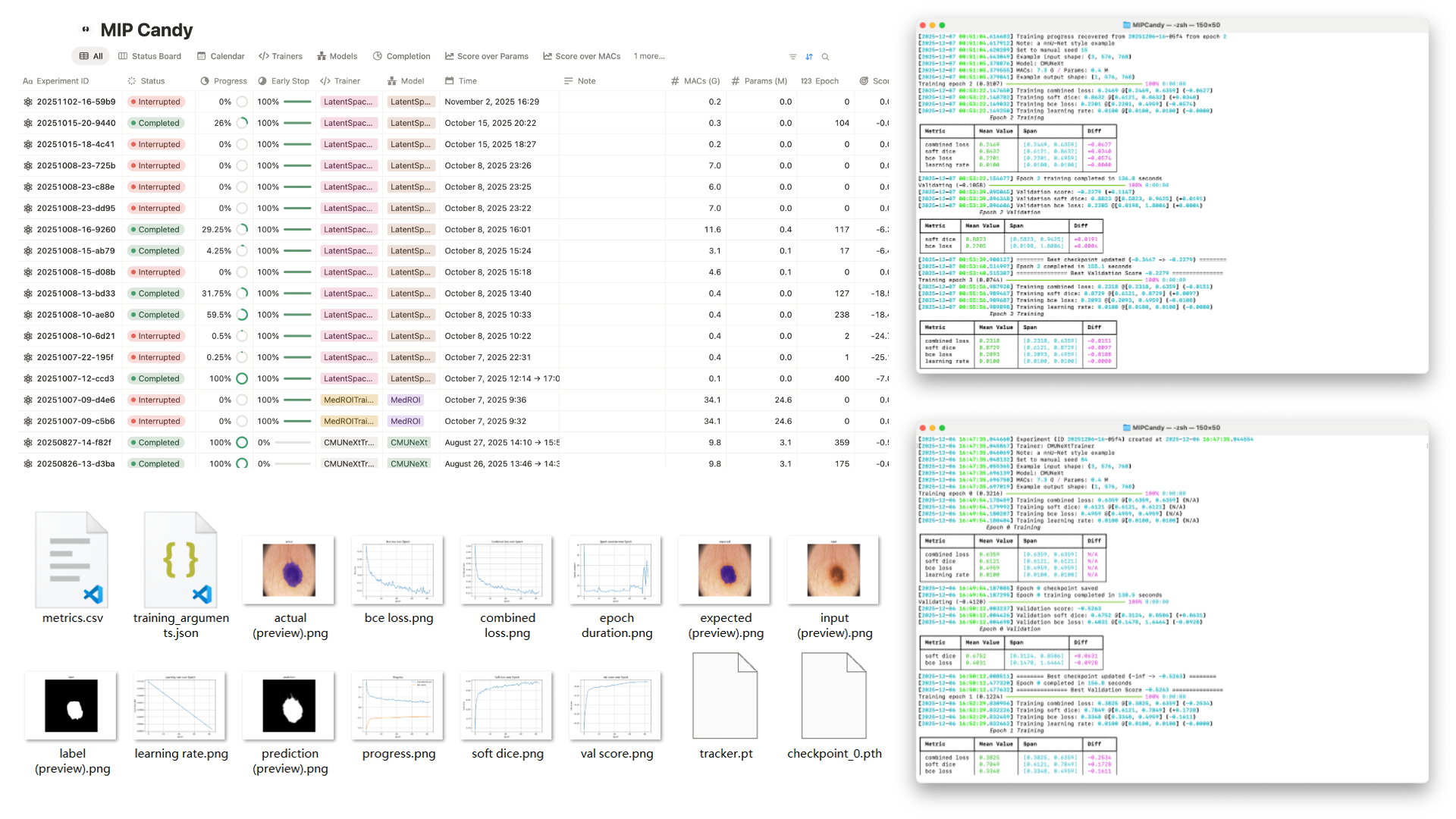
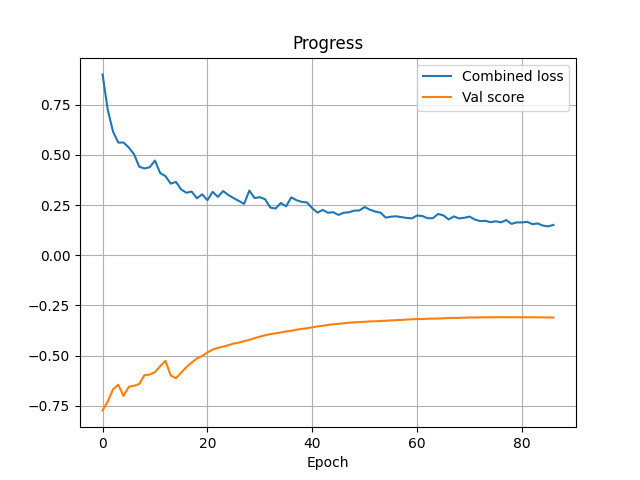
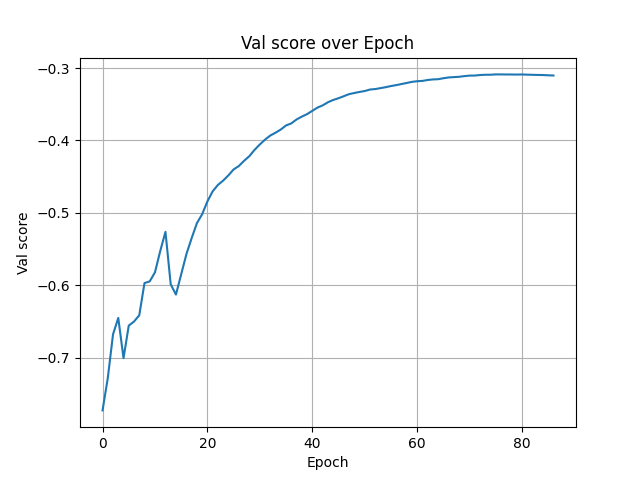
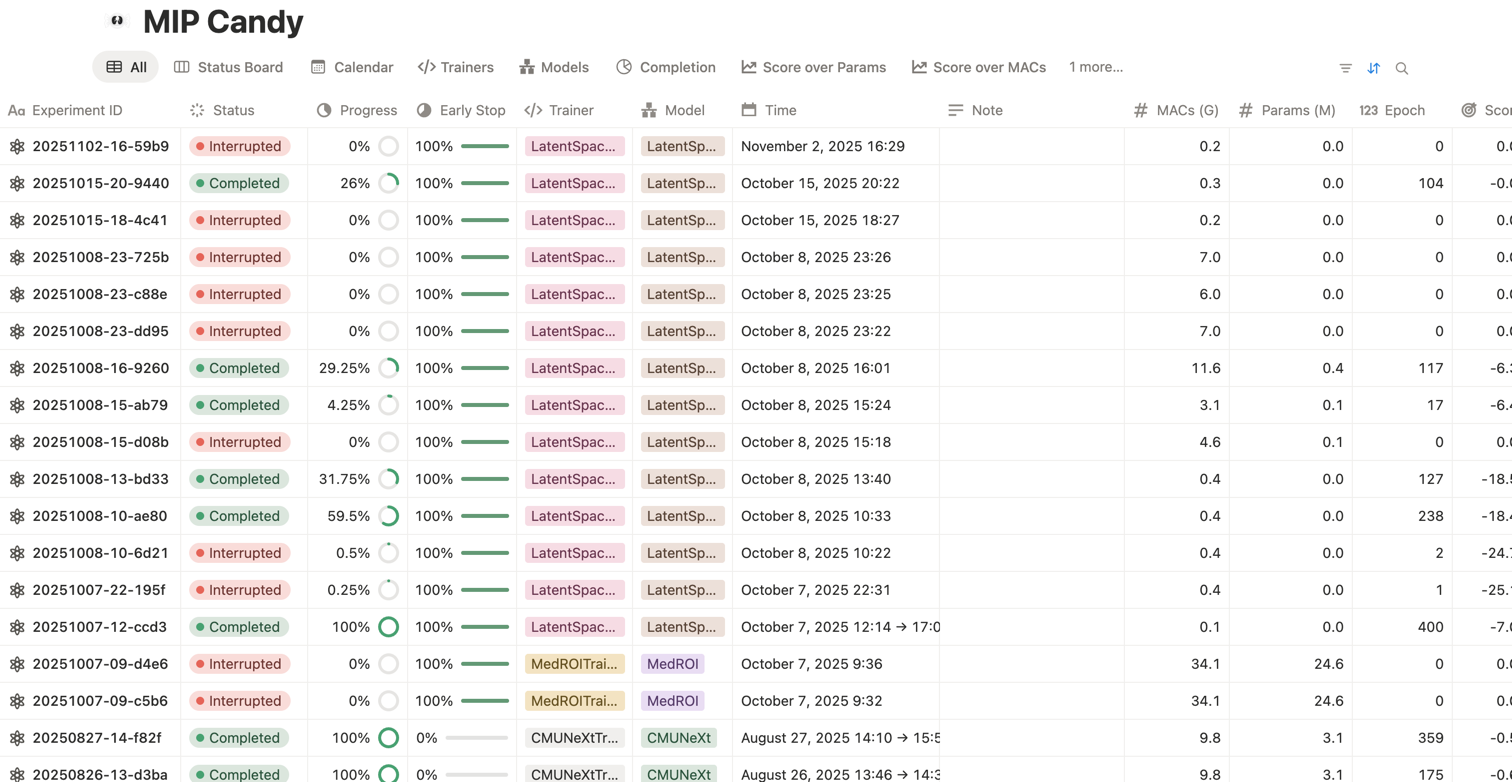
Quick start
Train like a Pro in a few lines
Download a dataset, create a dataset wrapper, and hand it to a trainer. Below is an example using
the ACDC dataset. The example code replicates most of nnU-Net's features but without augmentations.
Example: train on the ACDC dataset
from typing import override
from os.path import exists
from monai.networks.nets import DynUNet
from torch import nn
from torch.utils.data import DataLoader
from mipcandy import SegmentationTrainer, AmbiguousShape, auto_device, download_dataset, NNUNetDataset, inspect, \
load_inspection_annotations, RandomROIDataset
class UNetTrainer(SegmentationTrainer):
@override
def build_network(self, example_shape: AmbiguousShape) -> nn.Module:
kernel_size = [[3, 3, 3]] * 5
strides = [[1, 1, 1]] + [[2, 2, 2]] * 4
return DynUNet(spatial_dims=3, in_channels=example_shape[0], out_channels=self.num_classes,
kernel_size=kernel_size, strides=strides, upsample_kernel_size=strides,
deep_supervision=self.deep_supervision, deep_supr_num=2, res_block=True)
if __name__ == "__main__":
device = auto_device()
download_dataset("nnunet_datasets/ACDC", "tutorial/datasets/ACDC")
dataset = NNUNetDataset("tutorial/datasets/ACDC", align_spacing=True)
if exists("tutorial/datasets/ACDC/annotations.json"):
annotations = load_inspection_annotations("tutorial/datasets/ACDC/annotations.json", dataset)
else:
dataset.device(device=device)
annotations = inspect(dataset)
dataset.device(device="cpu")
annotations.save("tutorial/datasets/ACDC/annotations.json")
dataset = RandomROIDataset(annotations, 2)
train, val = dataset.fold()
train_loader = DataLoader(train, 2, True, num_workers=2, prefetch_factor=2, persistent_workers=True)
val_loader = DataLoader(val, 1, False)
trainer = UNetTrainer("tutorial", train_loader, val_loader, device=device)
trainer.train(1000, note="example with the ACDC dataset")